By: Lynn McCain
One of the most fascinating aspects of a career in cancer research is that one never knows when or where the next great discovery will occur. This was true of a recent breakthrough discovery made by the Dr. Russell Ryan laboratory at the University of Michigan Medical School.
Ryan, an Assistant Professor in the Department of Pathology and a member of the Rogel Cancer Center, was using a method called ChIP-Seq to map the regulatory elements that control gene expression in B-cell lymphomas. By analyzing the sequences found in those elements, Ryan’s research team hoped to deduce the identity of proteins called transcription factors that play key roles in activating cancer-causing genes. For comparison, they included a set of samples from B-cell acute lymphoblastic leukemia (B-ALL), a pediatric blood cancer that the group had not previously studied. They were shocked to find that active regulatory elements in B-ALL contained not only typical protein binding sequences but also simple repeats of the sequence “GGAA”, usually considered a form of “junk DNA” with no regulatory function.
“Many investigators ignore repeat elements in this sort of analysis”, said Ryan. “My bioinformatician decided not to filter them out, and it led us in a whole new direction.”
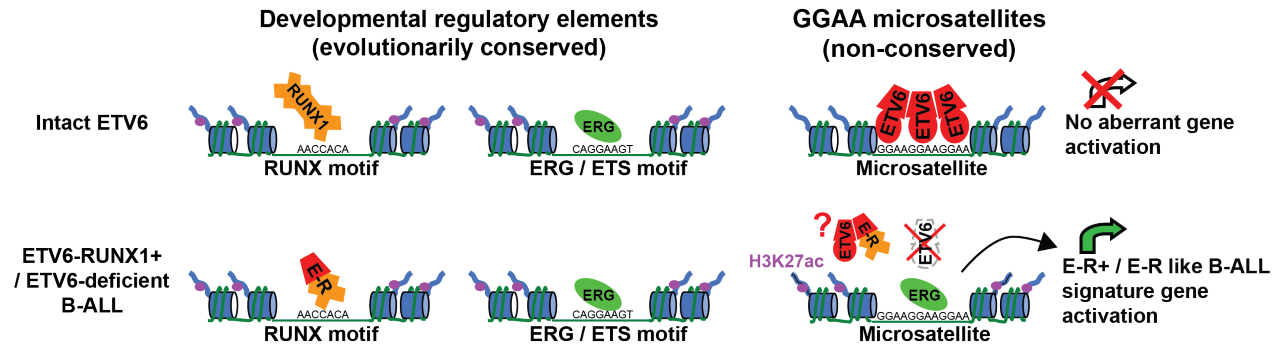
Ryan’s team soon realized that these ‘repeat enhancers’ were consistently active in a subtype of B-ALL called ETV6-RUNX1+ B-ALL, which is the second most common pediatric cancer in the United States, representing 25% of pediatric B-ALL cases. Although the defining oncogene of this type of leukemia, ETV6-RUNX1, itself encodes a transcription factor, Ryan’s team found that the ETV6-RUNX1 protein wasn’t directly activating the repeat enhancers. Instead, they found that the transcriptional repressor ETV6 normally binds to and silences these repeats. When the ETV6 gene is ‘broken’ by the formation of the ETV6-RUNX1 fusion, an activating protein called ERG replaces ETV6 at the repeats and converts them to active enhancers. By restoring ETV6 to the leukemia cells, Ryan’s team was able to shut down these repeat enhancers and identify the genes they activate, demonstrating that repeat enhancers control a known ‘gene expression signature’ of ETV6-RUNX1+ B-ALL. One particular repeat enhancer activated the EPOR gene whose expression is normally exclusive to red blood cells, allowing leukemia cells to respond to the growth-promoting blood hormone erythropoietin.
Ryan’s discovery appears to solve several longstanding mysteries. First, it suggests an explanation for why many ETV6-RUNX1+ leukemias gain additional mutations that destroy the second copy of the ETV6 gene. Second, it indicates why leukemias with other ETV6-disrupting mutations can often mimic the gene expression program of ETV6-RUNX1+ B-ALL. Finally, this discovery may explain why scientists have found it to be so difficult to re-create ETV6-RUNX1+ B-ALL in mouse models. While genes and normal regulatory elements are often well-conserved between humans and mice, “junk DNA” sequences like GGAA repeats are usually very different, meaning that repeat activation may not have the same effects in mice as they do in human leukemia.
The Ryan group’s discovery doesn’t yet indicate a new way to treat ETV6-RUNX1+ B-ALL, which is already curable in most cases after intensive chemotherapy, but that’s not the only way this new line of research might someday benefit patients. The first genetic events that lead to ETV6-RUNX1+ B-ALL occur in utero and are detectable in the blood of infants at birth, while other patients are born at high risk for leukemia due to pre-existing ETV6 mutations. By uncovering a fundamental epigenetic change that lies downstream of these key mutations, Ryan’s work may have taken us a step closer to predicting which of these patients will eventually develop leukemia, providing new opportunities for prevention or early intervention.
You can read more about this exciting research in Blood Cancer Discovery.
—
Citation:
Rohan Kodgule, Joshua W. Goldman, Alexander C. Monovich, Travis Saari, Athalee R. Aguilar, Cody N. Hall, Nihanka Rajesh, Juhi Gupta, Shi-Chun A. Chu, Li Ye, Ashwarya Gurumunthy, Ashwin Iyer, Noah A. Brown, Mark Y. Chang, Marcin P. Cieslik, Russell J. H. Ryan. ETV6 Deficiency Unlocks ERG-Dependent Microsatellite Enhancers to Drive Aberrant Gene Activation in B-Lymphoblastic Leukemia. Blood Cancer Discov (2023) 4 (1): 34–53.https://doi.org/10.1158/2643-3230.BCD-21-0224